Introducing camdl: engineering rigor for stochastic compartmental modelling
Disclosure: camdl is an open-source project I am developing at the Institute for Disease Modeling (Gates Foundation). I'm cross-posting here from internal channels since the project is public; the views expressed are my own and don't represent IDM or the Foundation.
Engineering scientific software is a different problem than general software engineering. The most consequential bugs in scientific code don’t crash loudly — they produce valid-looking results that get published in peer-reviewed papers. In 2006, Geoffrey Chang retracted five papers from Science, J. Mol. Biol., and PNAS because a sign-flip in his homemade data-reduction script had silently inverted the hand of his protein structures. Or take a favorite scientific bug example in genomics: more than a decade after Excel’s gene-symbol-to-date autoconversion was first documented , roughly a fifth of published supplementary tables still have it. And these are just the cases that got caught — the silent failures are by definition the ones that don’t.
The stakes of this default may be low when scientific code is just producing a figure for a paper. But in computational epidemiology, our software isn’t just producing an artifact, it’s informing decision-making in public health: outbreak responses, vaccination strategies, or major investments. The most consequential bugs in this domain aren’t crashes, they’re silently erroneous code that creates pernicious problems: trajectories that look plausible but lead decision makers astray — a per-capita rate written where the propensity needed a population-level one, a unit slip that survives review, uncertainty that should be conveyed but is silently dropped, a calibration that looks converged but isn’t. IDM’s pioneering agent-based simulator EMOD has a codebase that’s roughly 24% tests — a great early example of an engineering investment in scientific software long before it was standard. Eliminating entire classes of bugs should be, in my view, a central engineering goal of writing good epidemiological software.
A central challenge in the design of scientific software is the labor specialization of its users. An epidemiologist may spend a decade studying immunology, disease dynamics, vaccinology, etc., building domain expertise that is invaluable when it comes to producing good models. But such specialization has an unavoidable opportunity cost: every year spent on transmission dynamics is a year not spent on learning software engineering best practices, statistical inference algorithms, numerical analysis, compiler design, or software architecture. This problem is especially acute in low- and middle-income public-health offices, where research groups are often small and a single epidemiologist plays both modeller and software engineer — not for lack of capability, but for lack of the resources that wealthier research environments take for granted.
Since joining IDM, I’ve been curious whether domain-specific languages (DSLs)
could lower this engineering burden — letting modellers do rigorous inference
without first having to become rigorous software engineers. A DSL is a small
programming language purpose-built for one problem domain. The model’s code is
then just a declaration of the scientific model, written in a syntax that
reads like the whiteboard math you’d share with a colleague (e.g. infection : S --> I @ beta * S * I / N), and the runtime is the separate program that
actually executes it. A compiler1 sits between the DSL and the
runtime: it reads the model declaration, compiles it, checking extensively for
errors, and emits an intermediate representation (IR) the runtime consumes to
actually execute the simulation. With this DSL-compiler-IR architecture, the
modeller never needs to write the finicky, bug-prone code underneath — the
state-update loop, the propensity evaluator, the particle filter, the gradient
code. Instead, they describe the model; the compiler and runtime handle the
rest. Most scientific software tangles model and runtime implementation in one
script; the DSL-compiler-IR architecture keeps them apart. The closest
analogues are Stan
in statistics and
odin
for compartmental ODE modelling in R.
Both are excellent at what they do, and both heavily influenced camdl’s design:
Stan showed how far a probabilistic DSL with a serious compiler can go; odin
showed that a small, focused DSL can transform how a community writes its
models. camdl is in their lineage.2
But the DSL is only part of the answer. With camdl, I wanted the compiler doing more work to validate models — a goal that’s only gotten more urgent as coding agents start writing more model code (especially when the work is upstream of decision making). The compiler-level checks camdl does catch many mistakes regardless of who introduced them, human or agent; in both cases, the user gets friendly error, warning, or info messages. The compiler checks dimensions and units like a type system, differentiates rate expressions symbolically so inference gets exact gradients without an autodiff dependency, expands stratified models from a single declaration, and rules out subtle discretization bugs before anything runs.
I also wanted the runtime on the other side to match that effort: a full simulation and inference framework, with four simulation backends sharing one compiled model (Gillespie, tau-leaping, Euler-multinomial / chain-binomial, ODE) and backend compatibility checked before any step is taken, particle-filter methods that are standard for these inference problems (IF2 for MLE; PGAS+NUTS — a Gibbs split that updates latent trajectories with PGAS and parameters with NUTS — and PMMH for Bayesian posteriors), convergence gates that catch and flag bad model fits rather than passing them silently through, and content-addressable provenance so re-running a fit is a cache hit instead of a fresh run.
The compiler is written in OCaml (the same language Stan’s compiler uses, a fact I discovered only after I’d already made the same choice!), whose algebraic data types and pattern matching are what make serious compiler work tractable; the simulation runtime is in Rust, for memory safety and the performance this kind of engine needs.
Overall, camdl is my attempt to engineer a robust stochastic compartmental modeling framework around DSL architecture and cutting-edge statistical algorithms for model fitting. camdl isn’t a proof-of-concept: it has been technically validated by reproducing the findings of published disease models — He et al.’s 2010 fit of London’s measles data and the classic UK boarding school influenza outbreak — and ships with 1,204 tests, including continuous validation against pomp, scipy, and Kermack–McKendrick closed-form solutions, all run as CI gates that block merges on regression. The Garki Project Malaria Model and the Dhaka cholera dataset from King et al. (2008) are currently in flight as further external validations.
The rest of this post is a tour. I’ll start with what a real, fit-validated model looks like in camdl (the He et al. (2010) London measles SEIR, the canonical particle-filter benchmark) then walk through stratification at compile time, the compiler-level catches that rule out the silent bugs I described above, surveying the likelihood surface before committing to a fit, the scout / refine / validate calibration pipeline with its mandatory convergence gates, and the inference algorithms camdl ships.
A real model, written like the math
Here is a fragment of He, Ionides & King (2010) — the canonical particle-filter measles benchmark, fit to London weekly notifications 1950–1964, externally validated against pomp — written directly in camdl:
# He et al. (2010) London measles SEIR
# Full model: 138 lines — see he2010 vignette
forcing {
pop : interpolated { data = "covariates.tsv", value_col = population }
birthrate : interpolated { data = "covariates.tsv", value_col = birthrate }
school : periodic { period = 365.25 'days, step = 1 'days, on = [7:100, 115:199, 252:300, 308:356] }
}
let seas = 1.0 - amplitude + amplitude * (1.0 + 0.2411 / 0.7589) * school(t)
transitions {
infection : S --> E @ overdispersed(beta_base * seas * S * ((I + iota)^alpha) / pop(t), sigma_se)
latency : E --> I @ sigma * E
recovery : I --> R @ gamma * I
birth : --> S @ deterministic((1.0 - cohort) * birthrate(t) * pop(t) / 365.25)
# ... + per-compartment mu deaths
}
events {
cohort_entry : add(S, cohort * birthrate(t) * pop(t)) every 365.25 'days at_day 258
}
observations {
weekly_cases : {
projected = incidence(recovery)
every = 7 'days
likelihood = normal(mean = rho * projected,
sd = sqrt(rho * projected * (1 - rho + psi^2 * rho * projected)))
}
}Time-varying covariates, UK school-term forcing, overdispersed environmental noise on the force of infection, a once-per-year cohort pulse, and the He et al. heteroscedastic observation likelihood — all declarative, all compiler-checked. The full 138 lines of camdl (78 of actual code) build a model that pomp’s reference implementation expresses in 238 lines of mixed R, C Csnippet, and shell-wrapper glue — most of it plumbing rather than science:
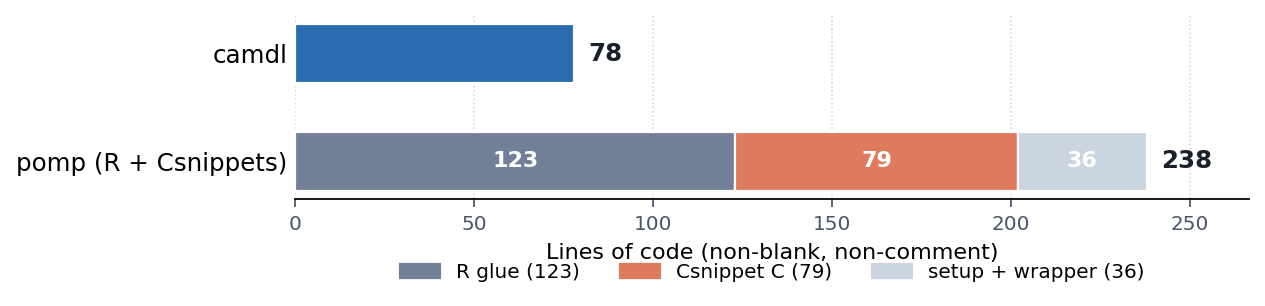
camdl owns the simulation engine, the propensity evaluator, and the inference stack; the modeller writes only the model. The responsibility split is sharp: the modeller is responsible for the model, and camdl is responsible for everything below the IR. If a compiling model produces a trajectory inconsistent with its declared semantics, that’s a camdl bug to find and fix.
Stratification is a compiler pass
The He et al. (2010) model above is one-dimensional — a single SEIR for all of London. But the models one may want to actually fit in policy work are almost never that simple: they’re stratified by age, by space, by risk group, by vaccination status. Hand-coding the stratified version is mostly mechanical bookkeeping (e.g. eight compartments instead of four, six transitions per age class instead of three, an age × age contact matrix to keep indexed correctly) but it’s roughly ten times the model code, and every line is a place for a typo to silently shift transmission across the wrong age class. In pomp this is hundreds of lines of C snippets; in NumPyro it’s vectorized indexing you write and own correctness for.
In camdl, stratification is something the compiler does for you. You
declare a dimensions {} block and a stratify directive, and the
compiler expands the base model into the full cartesian product before
anything else runs:
dimensions { age = [child, adult] }
stratify(by = age)
let N_local[a in age] = S[a] + E[a] + I[a] + R[a]
tables {
C_age : age × age = [[12.0, 4.0], [4.0, 8.0]] # contact matrix
}
transitions {
infection[a in age] : S[a] --> E[a]
@ beta * S[a] * sum(b in age, C_age[a, b] * I[b] / N_local[b])
progression[a in age] : E[a] --> I[a] @ sigma * E[a]
recovery[a in age] : I[a] --> R[a] @ gamma * I[a]
}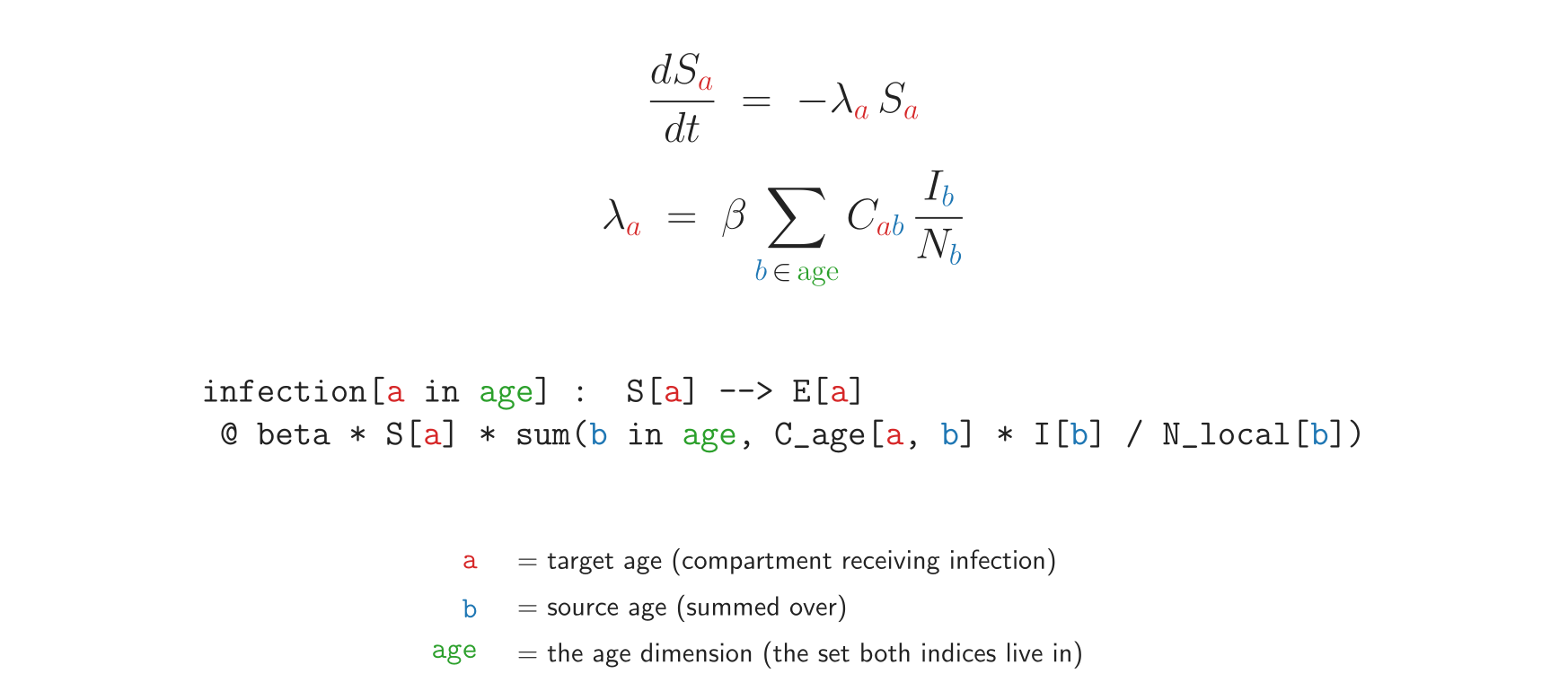
That’s the entire age-stratified SEIR. The compiler expands it to 8
compartments and 6 transitions, dim-checks the contact-matrix indexing
(a typoed dimension key is a compile error, not a silent zero), and
hands the runtime an IR that looks no different from the unstratified
case. Surveying, fitting, and held-out validation all work on the
expanded model unchanged. The same pattern composes: stratify(by = [age, space]) builds the full age × space product, with the same
guarantees.
A compiler doing what scripts can’t
The first version of camdl’s He et al. (2010) implementation carried a 12-nat
residual against pomp that had me scratching my head for two days. The cause was
a single line: (1 - exp(-rate * 1 'days)) — a propensity formulation that’s
correct only when the runtime integrator step happens to equal one day. Run it
at any other dt, and the fit silently calibrates to the wrong rate. Synthetic
recovery passed (the same bias appeared on both sides of the simulate/fit
loop), and no in-fit diagnostic surfaced it. The fix was a compiler lint
(L401) that catches the AST shape, points at the literal, and suggests the
dt primitive that makes the same expression dt-invariant. The bug would not
have surfaced without compiler-level inspection of the expression tree.
That’s the kind of bug camdl’s compiler is built to rule out. The most consequential modelling bugs are not crashes; they’re unit slips, dimension errors, and typos that produce trajectories that look plausible but are quietly wrong. The compiler catches each class before anything runs.
Units. Tick-prefixed annotations ('days, 'weeks, 'per_year)
are first-class; the compiler converts to the model’s time_unit.
When the grouping is ambiguous, it refuses rather than guessing:
$ camdl check bad_unit.camdl
error[E107]: ambiguous unit literal after '/': the unit suffix binds
to the adjacent number, not the whole expression. Use parentheses:
(20 / 100_000) 'per_year, or pre-compute: 0.0002 'per_year
┌─ bad_unit.camdl:9:17
│
9 │ let mu : rate = 20 / 100_000 'per_year
│ ~~~~~~~~~~~~~~~~~~~~~^Dimensions. Every expression carries an (P, T) exponent pair — counts are
(1, 0), per-capita rates are (0, -1), population-level rates are (1, -1).
The dim-checker propagates these through every operation and rejects
mismatches. This catches a very common modelling bug — writing a per-capita
rate where a total propensity is needed:
$ camdl check bad_dimension.camdl
error[E300]: transition 'infection' rate has wrong dimension
rate = ((beta * I) / (S + I + R))
expected dimension: P*T^-1 (population-level rate)
got dimension: T^-1 (per-capita rate)In pomp, NumPyro, or hand-rolled R, the second form compiles silently and produces trajectories where infection happens at the wrong rate.
Names. Typos that would silently introduce a new variable elsewhere become compile errors:
$ camdl check bad_name.camdl
error[E100]: undeclared name 'Q'
┌─ bad_name.camdl:11:33
│
11 │ recovery : I --> R @ gamma * Q
│ ^
= hint: check spelling, or add a declaration in
compartments/parameters/let/tablescamdl’s contribution is engineering rigor: typos, unit mismatches, and per-capita-vs-propensity confusion are exactly the class of bug a compiler can rule out, so scientific scrutiny can focus on the science.
A frame I keep coming back to here, borrowed from a personal interest in the Toyota Production System after reading Ohno’s Toyota Production System: Beyond Large-Scale Production : this is poka-yoke — error-proofing at the source — applied to the language scientific software is written in.
Surveying the likelihood surface before you fit
The hardest fitting bugs aren’t the ones where the chain fails to converge — those at least tell you something is wrong. They’re the ones where the chain converges confidently to the wrong answer, because the model has an identifiability problem you didn’t see coming: two parameters trading off along a ridge, a parameter pinned at a bound you set too tight, a region of parameter space the data simply cannot distinguish from another. Fitting a model you haven’t first surveyed is, in my experience, the single most common way to spend a week producing a result that doesn’t survive a careful look.
So before any camdl fit run, I almost always reach for camdl survey
— a diagnostic that maps the likelihood surface across the declared
parameter bounds:
$ camdl survey he2010_london.camdl --n-points 500 --renderIt draws 500 Latin-hypercube points across the parameter ranges (scale- aware, so log space for rates and linear for probabilities — the points actually span the parameter geometry instead of clumping near edges), evaluates the particle-filter loglik at each, and renders an interactive pair-plot HTML:

You see in one figure what would take a day of mis-spent IF2 runs to discover:
which parameters are well-identified, which trade off in ridges, where the
basin actually lives, and whether your bounds were sensible to begin with. The
brightest cluster of points is also the natural starting neighborhood for the
scout stage that comes next — and indeed, the scout can seed its IF2 chains
directly from the top-K survey points (init_method = "survey_top_k"). A
survey is a diagnostic, not a fitter; it doesn’t produce an MLE. But it’s the
cheapest hour of compute in the whole workflow, and it reliably saves many more
downstream.
Calibration is a workflow, not a script
Calibrating complicated epi models usually involves hand-assembled pipelines, which can easily grow in complexity and fragility as multiple models are compared. But with camdl, I wanted to explore ways to build canonical, safer, and more robust calibration pipelines as declarative artifacts:
# fit.toml — the entire calibration pipeline in one file
[model]
camdl = "he2010_london.camdl"
[data.observations]
weekly_cases = "data/cases.tsv"
[holdout]
weekly_cases = "data/cases_holdout.tsv" # held out, structurally unreachable
[estimate.R0] bounds = [10, 80]
[estimate.rho] bounds = [0.3, 0.6]
[estimate.sigma_se] bounds = [0.05, 0.20]
[stages.scout]
algorithm = "if2" # iterated filtering MLE
backend = "chain_binomial"
chains = 36
particles = 2000
iterations = 200
init_method = "lhs" # Latin-hypercube starts, scale-aware
[stages.refine]
algorithm = "pgas" # Bayesian posterior via PGAS+NUTS
backend = "chain_binomial"
starts_from = "scout" # seeds from scout's MLE
[stages.validate]
algorithm = "pfilter" # held-out predictive log-likelihood
particles = 4000
replicates = 8Three stages — scout, refine, validate — wired through one camdl fit run.
Between them sit convergence gates that fail loudly rather than just give
faulty results. Scout’s gate is two-legged: per-parameter chain agreement
(Gelman–Rubin-style on IF2’s per-iteration parameter means) plus a
decibans-spread check on chain-level log-likelihoods evaluated at high particle
count. Both legs must pass. Chains can agree on parameter values yet sit in a
basin tens to hundreds of decibans worse than truth — that’s the failure mode
the second leg catches. If either leg fails, the refine step refuses to start;
you fix the underlying issue (widen bounds, more chains, more iterations)
rather than passing a flag to ignore the gate.
A second gate runs after the fit finishes. The Richardson dt-convergence check evaluates the converged loglik on a halving ladder of integrator steps (dt, dt/2, dt/4) and refuses to bless a fit whose likelihood is still drifting. This is standard practice in numerical PDE work but is not, to my knowledge, automated in any epi-fit pipeline. The motivating bug was embarrassingly real: a plausible-looking boarding-school SIR fit reported a specific R₀ from a dt=1-day chain-binomial run. At dt=0.1, the same parameter vector scored 14 nats worse than the dt-converged MLE — the coarse step had created a phantom basin that the inference loop happily found, and no in-fit diagnostic at dt=1.0 could catch it (synthetic recovery passes, because the same dt on both sides cancels the bias). camdl now does this check automatically on every fit, emits a one-line PASS / MARGINAL / FAIL verdict, and saves the ladder to the run directory for inspection. The L401 lint and the Richardson check are complementary, not redundant: L401 catches the AST shape at compile time, and the Richardson check catches integration-step pathology after the fit, since it can arise even from dt-invariant rate expressions.
Returning to the TPS analogy: jidoka is the name for the other half of this discipline — stopping the line the moment an abnormality is detected, so the defect doesn’t reach the next station. Where poka-yoke makes the mistake impossible up front, jidoka catches it before it propagates; the compiler does the first, the gates do the second.
Moreover, camdl’s output is content-addressable. camdl fit run hashes the
full input vector — model IR, parameter values, seed, data files, algorithm
config, tool version — and writes to a directory keyed on that hash. An
unchanged config produces an unchanged hash, so a 2-day fit isn’t clobbered
by an accidental re-run; change one bound and the hash changes too, and a
fresh run starts without touching past results. camdl list enumerates every
fit the project has produced; camdl show <hash> surfaces the exact command
and seed; camdl cat <hash> emits the output. A camdl fit hash is effectively
a citation: paste it into a methods section and any reader with the source can
reproduce the result bit-for-bit. Reproducibility is a property of the tool,
not a discipline left to the user.
The inference stack
The fitting backend is not a wrapper around scipy; camdl ships the
methods you’d reach for in a serious paper, all driven from the same
fit.toml:
- Maximum likelihood via iterated filtering (IF2; Ionides et al., 2015 ) — fast point estimates and likelihood-surface mapping before committing to a full Bayesian run. Multi-start by Latin-hypercube within the parameter’s transform (log space for rates, linear for probabilities) so 16 chains span basins that 16 i.i.d.-uniform starts would clump in.
- Bayesian posteriors via PGAS+NUTS (Lindsten, Jordan & Schön,
2014
; Hoffman &
Gelman, 2014
) — honest
uncertainty on parameters and latent states, with the gradients NUTS
needs coming from a separate (and, in my view, underappreciated) place:
the compiler. The OCaml frontend walks each rate expression and emits
the analytical derivative as a source-to-source
rate_gradfield in the IR; the runtime evaluates exact ∇ log ℒ at the cost of one expression-tree walk per step. No autodiff tape, no finite differences, no JAX dependency. The expression language is first-order and pure — the same property that makes the dim-checker tractable also makes symbolic differentiation a 30-line OCaml pattern match. - Gradient-free fallback via PMMH (Andrieu, Doucet & Holenstein, 2010 ) — for posterior geometries where NUTS struggles, or when you want a second sampler.
- Profile likelihoods (Raue et al., 2009 ) — expose identifiability problems before you’ve spent months building around unidentifiable parameters.
- Out-of-sample validation via prequential scoring (Dawid, 1984 ; Gneiting & Raftery, 2007 ) — held-out predictive performance is the only honest test of fit, and the only honest basis for model comparison.
When you actually want to choose a model, camdl compare gives you
the decision in one view:
$ camdl compare preq_poisson.json preq_negbin.json preq_procnoise.json
Model T_score elpd Δelpd se(Δ) evidence PIT_cov90
─────────────────────────────────────────────────────────────────────────────────────
preq_poisson.json 14 -62.75 -7.44 5.52 -32.3 dB, decisive against 0.71
preq_negbin.json 14 -56.44 -1.12 1.31 -4.9 dB, indeterminate 0.93
preq_procnoise.json 14 -55.32 — — — 0.93
Δelpd is shown relative to the best-supported model (at the bottom); a more-negative Δelpd means worse predictive performance.Three columns to read. Δelpd is the expected-log-pointwise-predictive- density gap between this model and the best-supported one (zero at the bottom row by construction; more-negative means worse held-out prediction). The evidence column converts that gap to decibans (10 × log₁₀ of the likelihood ratio, so −10 dB means the data favour the other model by 10:1; see Kass & Raftery 1995 for the conventional “weak / positive / strong / decisive” scale). Probability-Integral-Transform (PIT) coverage at 90% is the fraction of held-out observations whose true value falls inside the model’s 90% predictive interval — near 0.90 for a well-calibrated model. All three columns are computed from held-out predictive scoring of the same boarding-school epidemic data fit under three observation models. The 0.71 PIT coverage on Poisson flags it as underdispersed; NegBin and the process-noise variant are both well-calibrated at 0.93. The baseline pivots automatically to the best-supported model. This is the kind of output that makes routine model comparison cheap enough to actually do.
The credibility anchor: where camdl overlaps with established reference
implementations, camdl matches — and not just statistically. The periodic-B-spline
forcing the He et al. (2010) chapter uses agrees with both pomp 6.4 (R) and
scipy.interpolate.BSpline (Python) at less than 10⁻¹² max absolute difference
across a 200-point grid. Two independent ecosystems with non-overlapping code
paths, both bit-identical. The Fourier forcing matches numpy to the same
tolerance. The He et al. (2010) London measles MLE recovers pomp’s published
values within particle-filter Monte Carlo error; the boarding-school SIR
matches pomp’s tutorial within statistical tolerance; the bare SIR final-size
hits the Kermack–McKendrick closed form. Every release runs these as CI gates
— not as one-off validation studies but as continuous tests that block merges
if camdl ever drifts.
Why a DSL pays off here
Compartmental + camdl is often structurally cheaper than ABMs for the
calibration questions most epi modellers face. ABMs scale with population ×
steps × interactions, and their calibration scales worse — bespoke ABC,
summary-statistic matching, no off-the-shelf likelihoods. By contrast, camdl
scales with parameter count and observation richness; you get full likelihoods,
gradients, posteriors, profile identifiability checks, and held-out predictive
scoring out of the box. For policy questions where you don’t need agent
heterogeneity, the cost asymmetry is one to two orders of magnitude on
prototyping — cheaper model writing means more variants can be tried in the
same wallclock time, and camdl compare makes the head-to-head quantitative
rather than asserted.
The DSL is what unlocks this. Three concrete leverage points.
Small files, full fidelity. The He et al. (2010) model takes 78 lines of camdl versus 238 lines of pomp R + Csnippets + wrapper for the same model. Most of the camdl file is the science; most of the pomp file is plumbing.
Compiler errors that LLMs can act on. The same properties that catch human bugs — small declarative files, structured error codes (
E100,E300), no implementation glue — make camdl LLM-friendly by construction. An agent readingerror[E300]: expected dimension P·T⁻¹, got T⁻¹knows exactly what’s wrong and how to fix it; a typical compartmental model fits comfortably in a single context window. In my experience, Claude Code often one-shots a working model from a short prompt (“build me an SEIR with age structure and weekly NegBin observations”), or converges in a round or two against the compiler when it doesn’t. The compiler is the ground truth; the agent doesn’t have to guess what a “rate” means in this codebase.Engineering rigor without an engineering team. A small team in an LMIC public-health office, a single-modeler academic group, a postdoc fitting a paper on a deadline — none of these have the bandwidth to maintain a custom calibration pipeline that tracks upstream pomp/Stan changes, validates against published references, and enforces convergence diagnostics on every fit. camdl’s compiler does that work. The fit pipeline is one TOML file. The convergence gates are mandatory. The provenance is content-addressable. The same infrastructure that makes camdl LLM-friendly makes it accessible to modellers without a software-engineering substrate behind them.
On AI-assisted scientific software
camdl reached this alpha in 974 commits over nine weeks; my first commit was on March 12. I like to do little back-of-the-envelope type calculations to anchor things in units that make relative sense. So to anchor the pace concretely: the repo is currently ~167,000 lines and ~6.85 MB of source text across OCaml, Rust, Python, Shell, Markdown, and config.
I type about 74 wpm on my laptop keyboard, which is about 22,200 characters per hour. If I personally were to type out the entire repo at my max typing speed, eight hours a day, five days a week, with zero pauses to think, debug, eat, or sleep, it would take ~7.7 work-weeks. The actual elapsed time was ~9 weeks. So even if I had done nothing but transcribe pre-written code at full speed for every working hour of the project, I would have just barely fit it into the calendar — with no time left over for design, debugging, statistical work, or any of the actual engineering. (To be fair, the version of those nine weeks involving no emails, no meetings, and no Teams does have a certain monastic appeal.) At a realistic professional rate for compiler-and-Rust development — somewhere in the 10–50 finished-LoC-per-hour range, since that metric is famously noisy — solo hand-writing this codebase would have taken on the order of 1 to 3 years. When there’s less than twenty years left , we can’t spend 1 to 3 of them just building an easy-to-use, AI-friendly compartmental-modelling DSL — we need to be using it.
So the pace isn’t possible with hand-typing alone, and it raises a question worth answering directly: how do you square AI-assisted development with the engineering rigor this post has been arguing for?
Honestly, we don’t yet know. The space of AI-assisted scientific software is too new to have settled norms, and what counts as adequate oversight is now, in many ways, an open empirical question. What I can say is what I tried to do: rapidly de-risk the DSL-with-serious-compiler architecture for stochastic compartmental modelling, and at the same time build a tool that made writing and fitting compartmental models easier (and hopefully fun).
In the last eight months or so I have seen agentic coding tools get incredibly good at writing scientific software, bashing out intricate numerical algorithms carefully and in far less time than I could.3 The bet I made on the engineering side is that lots of planning iteration, compiled languages with algebraic types, thinking in terms of types (and lots of consolidation refactoring), and 1,204 tests would produce a software product with about the same number of bugs as the average alpha release — and hopefully fewer. The same compiler-checked everything and CI gates that make camdl trustworthy for users are the infrastructure that keeps AI-assisted development in the rigorous mode rather than the vibe-coded one: when the agent gets something wrong, it shows up as a compile error or a failing test, not as a silent bug.
My hope is that this AI-driven workflow I’ve been figuring out was enough to sculpt Claude into something like a second-year grad student that’s really good at coding, overseen in regular meetings by a graduate advisor. I think that’s about where we’re at with the interaction between scientists and AI.
Whether that bet holds up is exactly the kind of thing the field should investigate openly. I actively welcome readers to stress-test this codebase — finding bugs, studying where the agent and I converged on bad design, characterizing what kinds of prompting or test infrastructure would have caught what. The questions about whether AI assistance preserves rigor at this scale matter beyond just camdl, and the work matters to public health.
This generalizes back to the user side, which is where I think it matters most. A scientist using camdl with Anthropic’s Claude Code — the same daily collaborator that made this project possible at this pace — gets the same arrangement I had as the developer: the compiler catches what the agent gets wrong, the convergence gates catch what the inference gets wrong, and the modeller is the one whose judgment determines whether the model is the right model for the question. AI is leverage; the standards belong to the person.
A specific thanks here to the many, many, Claude Code agents that served me along the way: through these nine weeks Claude has been a daily pair-programmer in OCaml and Rust, a sanity-checker on the statistical derivations, a quick summarizer of methods papers, a partner in debugging the gnarlier corners of the inference stack, and the thing that held a multi-layer codebase in working memory while I focused on any one piece of it. The scientific judgment was mostly mine; the design was something we wove together; the bugs were mostly Claude’s authorship and mine to chase (though I’d have produced a comparable litany on my own — probably more). The engineering tempo this post represents reflects that collaboration.
Try it
This blog post marks camdl’s alpha release — usable for real fits, with a stable enough public surface that we’re documenting it openly, but with breaking changes still expected. If you want to start:
- Read the documentation/vignettes. vsbuffalo.github.io/camdl-docs walks through everything above with executable examples; Getting Started is the entry point.
Send feedback. Source is at github.com/vsbuffalo/camdl . What I most want to hear at alpha: where the error messages are unhelpful, what camdl language features are missing or feel clunky, where the docs are wrong, what frictions exist during model building and fitting, and which published models you’d want to see reproduced next.
Contribute. camdl is far from feature-complete. Code contributions, model ports, runtime-backend extensions, and stress-testing on real models that surfaces bugs are all welcome. There’s a long backlog of features I’d like camdl to grow into — open an issue or get in touch and I can point you at something matched to your interests.
I’ve had a background interest in compilers and interpreters for a while, since as an undergrad I worked with UC Davis professor and R-core contributor Duncan Temple Lang on RLLVMCompile , an experimental LLVM-based compiler for R. I also caught a brief case of the Lisp bug from Paul Graham’s essays, and wrote a small Lisp interpreter. ↩︎
IDM also has prior art in this genre: CMS , a cool Lisp-like S-expression DSL for compartmental models with several excellent backend solver implementations. I came across it only after starting camdl, so it didn’t shape the design — but it’s the closest in-house precedent and worth knowing about. ↩︎
Though I do miss the sweet, sweet payoff that comes after a week of head-bashing over a numerical underflow, a sign error in the gradient, or a fixed-point iteration that won’t converge — when you finally find the one missing log-sum-exp and the traces snap into agreement with the analytic solution. AI-assisted work does, honestly, trade some of that away. ↩︎